Dortmund researchers Max Planck unveils the smallest details of actin filaments
Actin filaments are protein fibers that make up the internal structure of the cell. As active elements in our cells, they support cell fusion and cell movement and are involved in many other cellular processes. They’re also an important key component of muscle cells, and researchers at the Max Planck Institute for Molecular Physiology in Dortmund have succeeded for the first time in making hundreds of water molecules visible in actin filaments – a quantum leap in actin research. With the help of cryo-electron microscopy (cryo-EM), Stefan Raunser’s group shows in unprecedented detail how actin proteins are arranged into filaments, how ATP – the cell’s energy source – is located in the protein pocket and where individual water molecules position and interact with ATP.
© MPI Molecular Physiology
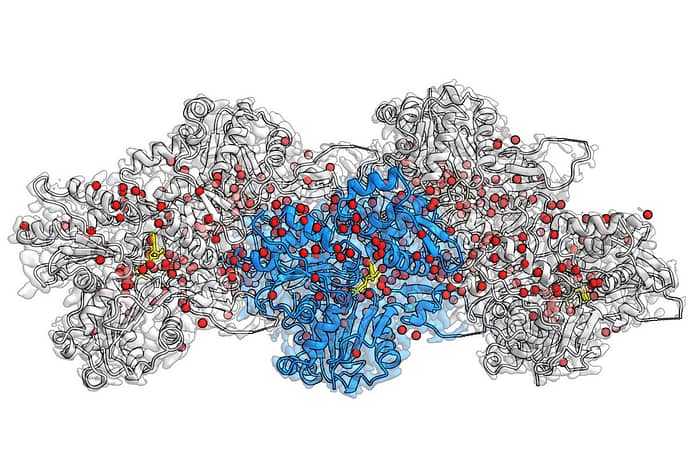
Cryo-EM reconstruction of F-actin bound to Mg2+-ADP-BeF3– It is resolved with 2.2 Å. The central actin subunit is colored blue, the other four subunits are grey. The corresponding densities of water molecules are colored red. ADP is yellow. © MPI Molecular Physiology
“We answer fundamental questions in life that researchers have been trying to answer for decades,” says Raunser. In eukaryotic cells, actin proteins are abundant and assembled into filaments. A network of these filaments forms the cell’s cytoskeleton, which controls various cellular processes through locomotion. So use, for example, immune cells use actin filaments to move and hunt down bacteria and viruses. It was previously known that the dynamics of the filaments is regulated by ATP hydrolysis – the reaction of ATP with water, in which the phosphate group is split and energy is produced. However, the molecular details behind this process remain unclear so far.
Because actin filaments are too flexible or too large for X-ray and NMR crystallization, detailed images of the filaments can only be obtained with cryo-EM. In 2015, Raunser’s team used this technology to create a new 3D atomic model of the filament with a resolution of 0.37 nanometers. In 2018, his group described the three different states assumed by the actin protein in filaments: ATP-bound, ADP-bound in the presence of cleaved phosphate, and ADP-bound after phosphate release.
How do water molecules move?
In their current study, Stefan Raunser’s team was able to set a new resolution record: they obtained structures for all three actin states with a resolution of ~0.2 nanometers and thus were able to make previously invisible details visible. The 3D maps not only show all the amino acid side chains of the proteins, but also reveal where hundreds of water molecules are located.
By comparing these new structures with those of the isolated actin protein, they were able to infer how the water molecules move. During polymerization, the water molecules shift in the ATP pocket so that only one water molecule remains in front of the ATP, ready to attack the phosphate and start hydrolysis. The precise insights gained through this approach have the potential to advance further research in this area: “Our high-resolution model could support researchers in developing small molecules for light microscopy on tissues and ultimately for therapeutic applications,” says Raunsir.
Did you open a door?
The authors also shed light on the fate of phosphates. Until now, researchers believed that there was a back door in the ATP pocket that remained open after ATP hydrolysis to facilitate phosphate exit. However, the new cryo-EM structures do not show an open back door. Therefore, the mechanism of phosphate release remains a mystery. “We think there really is a door. But it probably only opens for a while and then closes again,” comments Raunser, who now wants to use mathematical simulations and time-resolved electromagnetic cooling methods to show how the phosphates exit. Clearly, these exciting discoveries opened the door for researchers to discover more details about the processes by which actin filaments contribute to cell movement.